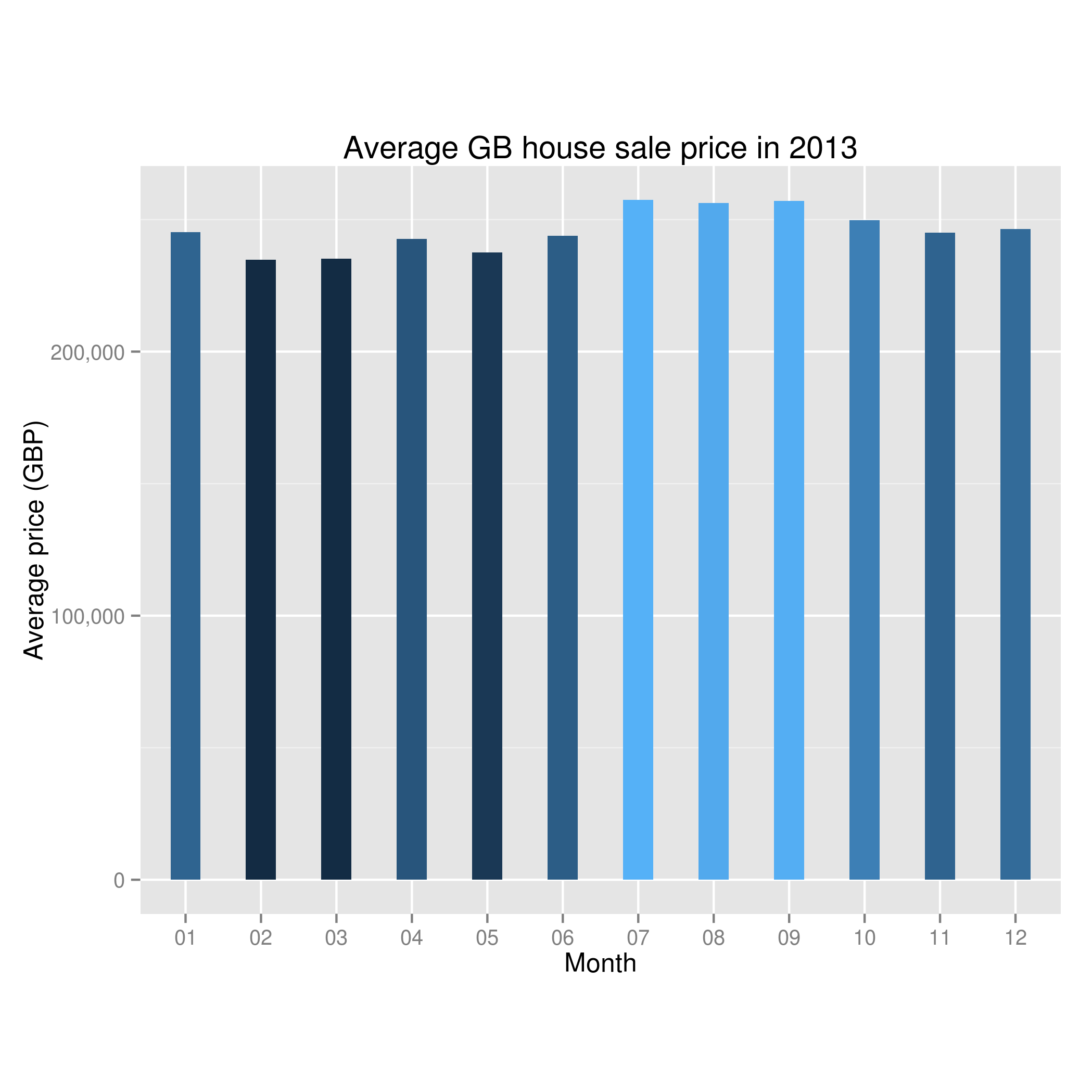
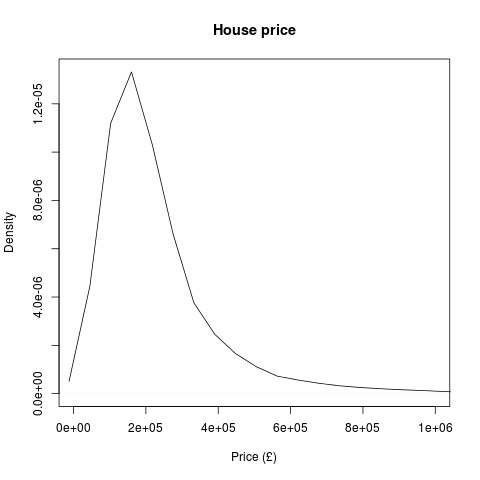
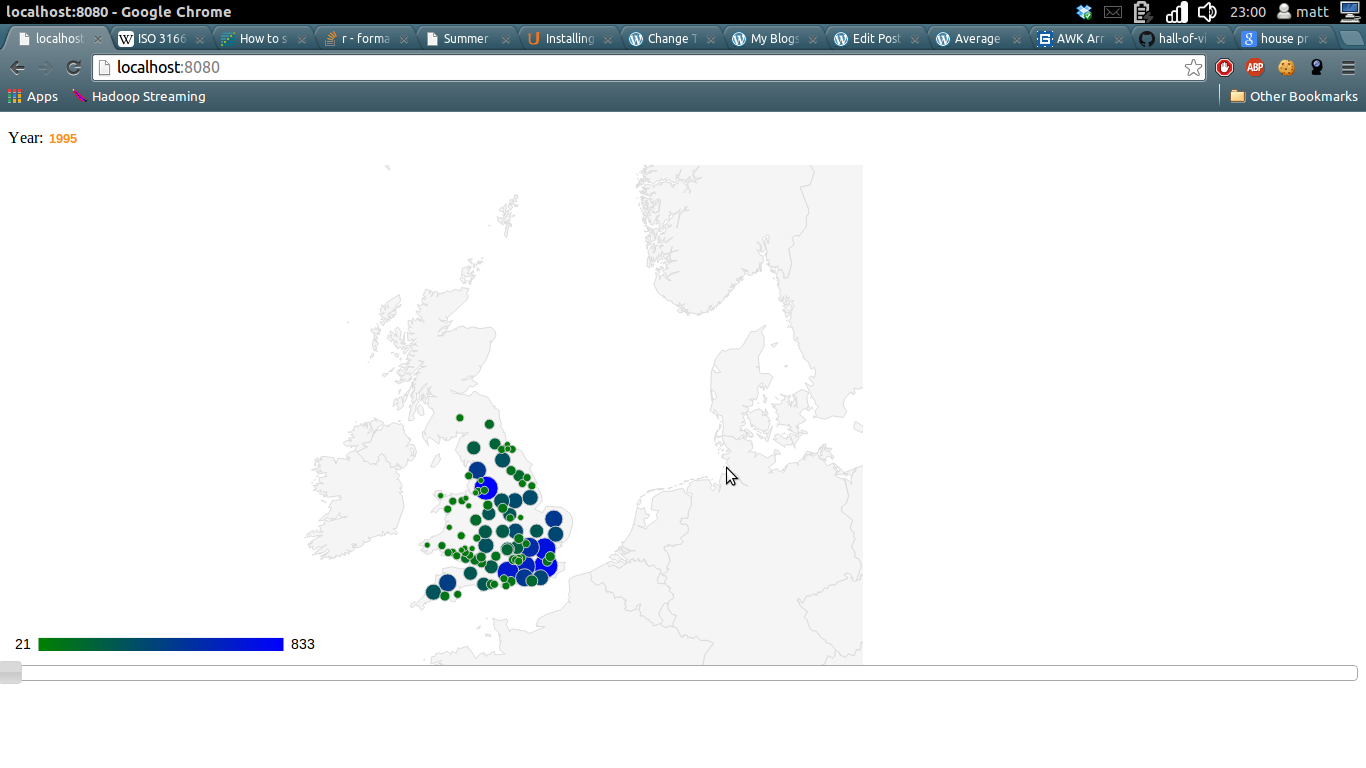
A Data Scientist happened upon a load of stuff – junk, at first glance – and wondered, as was his wont, “what can I get out of this?”
Conceded: as an opening line this is less suited to a tech blog than an old-fashioned yarn. In an (arguably) funny way, this isn’t far from the truth. My answer: get some holism down your neck. Make it into a modest, non-production Hadoop cluster and enjoy a large amount of fault-tolerant storage, faster processing of large files than you’d get on a single high-spec machine, the safety of not having placed all your data-eggs in one basket, and an interesting challenge. Squeeze the final, and not inconsiderable, bit of business value out of it.
To explain, when I say “stuff”, what I mean is 6 reasonable but no longer DC-standard rack servers, and more discarded dev desktops than you can shake a duster at. News of the former reached my Data Scientist colleague and I by way of a last call before they went into the skip; I found the latter buried in the boiler room when looking for bonus cabling. As a northerner with a correspondingly traumatic upbringing, instinct won out and, being unable to see it thrown away, I requested to use the hardware.
I’m not gonna lie. They were literally dumped at my feet, “unsupported”. Fortunately, the same qualities of character that refused to see the computers go to waste saw me through the backbreaking physical labour of racking and cabling them up. Having installed Ubuntu Server 13 on each of the boxes, I had soon pinged my desktop upstairs successfully and could flee the freezing server room to administrate from upstairs. Things picked up from here, generally speaking.
The hurdle immediately ahead was the formality of installing and correctly configuring Hadoop on all of the boxes, and this, you may be glad to know, brings me to the point of this blog post. Those making their first tentative steps into the world of Hadoop may be interested to know how exactly this was achieved, and indeed, I defy anyone to point me towards a comprehensive Hadoop-from-scratch quick start which leaves you with a working version of a recent release of Hadoop. Were it not for the fact that Hadoop 2.x has significant configuration differences to Hadoop 1.x, Michael Noll’s excellently put-together page would be ideal. It’s still a superb pointer in itself and was valuable to me during my first youthful fumblings with Hadoop 18 months ago. The inclusion of important lines of bash neatly quashes the sorts of ambiguity that may arise from instructions like “move the file to HDFS” which you sometimes find.
In any case, motivated by the keenness to see cool technology adopted as easily and widely as possible, I propose in this to briefly explain the configuration steps necessary to get me into a state of reverse cartography. (Acknowledged irony: there will probably be a time when someone reads this and it’s out of date. Apologies in advance.) Having set up a single node, it’s actually more of a hassle to backtrack over your configuration to add more nodes than to just go straight to a multi-node cluster. Here’s now to do the latter.
Setting the Scene
The Hadoop architecture can be summarised in saying that it elegantly facilitates doing two things in a distributed manner: storing files, and processing files. The two poles of the Hadoop world which respectively deal with these are known as the DFS (distributed file system) layer, and the MapReduce layer. Each layer knows about the other, but can, and indeed must, be configured, administrated and used fairly independently across a cluster. There’s an interesting history to both of these computing paradigms, and many papers written by the likes of Google and Facebook describing their inner workings. A quick Youtube yields some equally illuminating talks. My personal favourites on the two layers are this for HDFS and this for MapReduce.
Typically a cluster of n computers (nodes) will have 1 master node, which coordinates the distribution of storage and processing, and (n-1) slave nodes which do the actual stuff. The modules (daemons) in Hadoop 2.x which control all of these are as below.
|
Master |
Slaves |
| DFS |
Namenode |
Datanode |
| MR |
Resourcemanager |
Nodemanager |
Obligatory diagram:
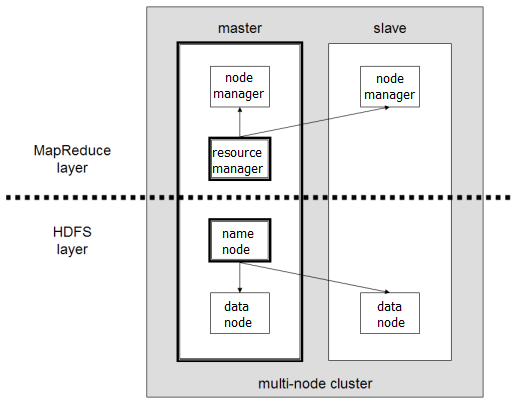
Illuminating.
Based on your current cluster setup, Hadoop makes a bunch of intelligent decisions about where to put files, and which machines to do certain bits of processing on, motivated by maximising redundancy and fault tolerance by clever replication choices, minimising network overhead, optimally leveraging each bit of hardware, and so on. The way that the architecture makes these decisions in such a way that you, the Hadoop developer, don’t have to worry about them, is where the real beauty and power of Hadoop lies. We’ll see later in this blog how, whilst HDFS and MapReduce are breathtakingly complex and scalable under the bonnet, leveraging their power is no more difficult than performing normal straightforward file system operations and file processing in Linux.
So. At this stage, all you have is a collection of virginal, disparate machines that can see each other on the network, but beyond that share no particular sense of togetherness. Each must undergo the same setup procedure before it’s ready to pull its weight in the cluster. In a production environment, this would be achieved by means of an automated deployment script, so that nodes could be added easily and arbitrarily, but that is both overkill and an unnecessary complication here. Good old-fasioned Bash elbow grease will see us through.
Having said that, one expedient whose virtues I will extol is a little gem of software called SuperPutty, which will send the same command from any single Windows PC to all the Linux boxes simultaneously, in so doing greatly reducing repetitiveness and cutting out chances for human error:
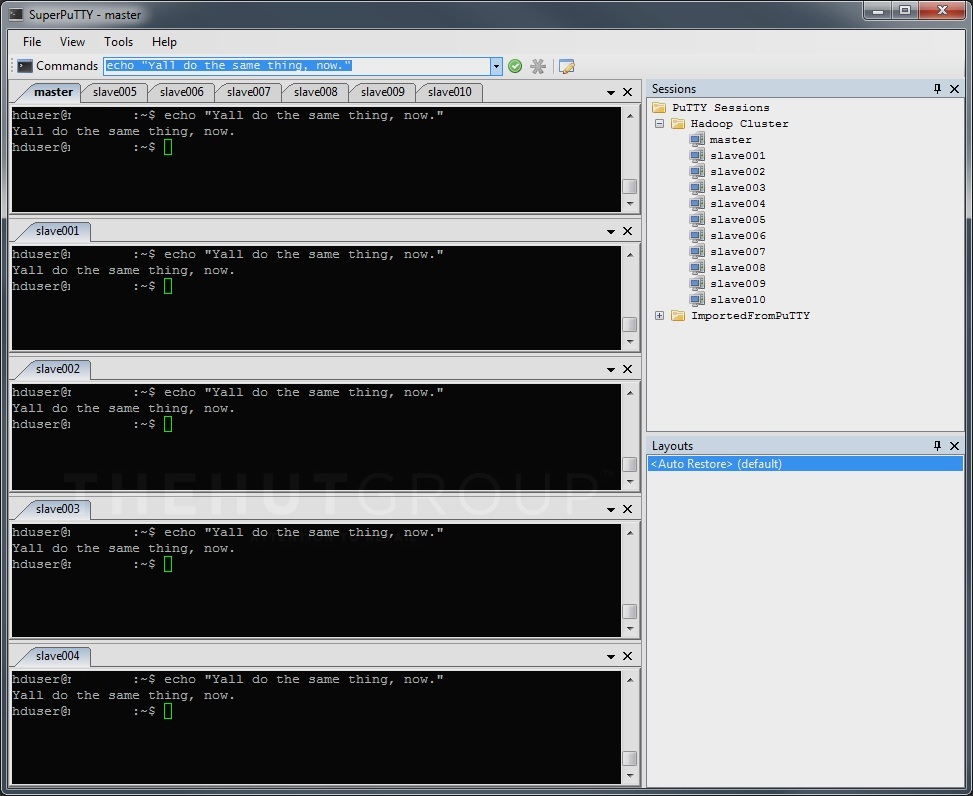
Using SuperPutty to send commands en-masse is only the same as doing the same thing on each box in sequence.
Connect to all the boxes and make sure you’re at the same bash prompt on all of them. SuperPutty will let you store connection authentication details to save you even more time in swiftly connecting to every machine in your cluster. (Disclaimer: if you do store passwords, anyone with Linux knowledge who finds your unattended, unlocked PC could connect to your cluster and perform wild-rogue Hadoop operations on your data. Think carefully.)
Masters and Slaves
One of your computers will be the master node, and the rest slaves. The master’s disks are the only ones that need to have an appropriate RAID configuration, since Hadoop itself handles replication in a better way in HDFS: choose JBOD for the slaves. If one of your machines stands above the rest in terms of RAM and/or processing power, choose this as the master.
Since Hadoop juggles data around amongst nodes like there’s no tomorrow, there are a few networking prerequisites to sort, to make sure it can do this unimpeded and all nodes can communicate freely with each other.
Hosts
Working with IPs is a lot like teaching cats to read: it quickly becomes tedious. The file /etc/hosts enables you to specify names for IP addresses, then you can just use the names. Every node needs to know about every other node. You’ll want your hosts file on each of the boxes to look something like this so you can refer to (eg) slave 11 without having to know (or calculate!) slave 11’s IP:
123.1.1.25 master
123.1.1.26 slave001
123.1.1.27 slave002
123.1.1.28 slave003
123.1.1.29 slave004
... etc
It’s also a good idea to disable IPv6 on the Hadoop boxes to avoid potential confusion regarding localhost addresses… Fire every box the below commands to append the necessary lines to /etc/sysctl.conf…
sean@node:~$ echo "#disable ipv6" | sudo tee -a /etc/sysctl.conf
sean@node:~$ echo "net.ipv6.conf.all.disable_ipv6 = 1" | sudo tee -a /etc/sysctl.conf
sean@node:~$ echo "net.ipv6.conf.default.disable_ipv6 = 1" | sudo tee -a /etc/sysctl.conf
sean@node:~$ echo "net.ipv6.conf.lo.disable_ipv6 = 1" | sudo tee -a /etc/sysctl.conf
The machines need to be rebooted for the changes to come into effect…
sean@node:~$ sudo shutdown -r now
Once they come back up, run the following to check whether IPv6 has indeed been disabled. A value of 1 would indicate that all is well.
sean@node:~$ cat /proc/sys/net/ipv6/conf/all/disable_ipv6
Setting up the Hadoop User
For uniformity across your cluster, you’ll want to have a dedicated Hadoop user with which to connect and do work…
sean@node:~$ sudo addgroup hadoop
sean@node:~$ sudo adduser --ingroup hadoop hduser
sean@node:~$ sudo adduser hduser sudo
We’ll now switch users and work as the new Hadoop user…
sean@node:~$ su - hduser
hduser@node:~$
SSH Promiscuity
Communication between nodes take place by way of the secure shell (SSH) protocol. The idea is to enable every box to passwordlessly use an SSH connection to itself, and then copy those authentication details to every other box in the cluster, so that any given box is on familiar terms with any other and Hadoop is unshackled to work its magic!
Firstly, send every box the instruction to make a passwordless SSH key to itself for hduser:
hduser@node:~$ ssh-keygen -t rsa -P ""
Bash will prompt you for a location in which to store this newly-created key. Just press enter for default:
Generating public/private rsa key pair.
Enter file in which to save the key (/home/hduser/.ssh/id_rsa): Created directory '/home/hduser/.ssh'.
Your identification has been saved in /home/hduser/.ssh/id_rsa.
Your public key has been saved in /home/hduser/.ssh/id_rsa.pub
The key fingerprint is: 9b:82...................:0e:d2 hduser@ubuntu
The key's randomart image is: [weird ascii image]
Copy this new key into the local list of authorised keys:
hduser@node:~$ cat $HOME/.ssh/id_rsa.pub >> $HOME/.ssh/authorized_keys
The final step in enabling local SSH is to connect – this will save the fingerprint of the host to the list of familiar hosts.
hduser@node:~$ ssh hduser@localhost
The authenticity of host 'localhost (::1)' can't be established.
RSA key fingerprint is d7:87...............:36:26
Are you sure you want to continue connecting? yes
Warning: permanently added 'localhost' (RSA) to the list of known hosts.
Now, to allow all the boxes to enjoy the same level of familiarity with each other, fire them all this command:
hduser@node:~$ ssh-copy-id -i $HOME/.ssh/id_rsa.pub hduser@master
This will make every box send its SSH key to the master node. Unfortunately, you have to repeat this to tell every box to send its key to every node…
hduser@node:~$ ssh-copy-id -i $HOME/.ssh/id_rsa.pub hduser@slave001
hduser@node:~$ ssh-copy-id -i $HOME/.ssh/id_rsa.pub hduser@slave002
hduser@node:~$ ssh-copy-id -i $HOME/.ssh/id_rsa.pub hduser@slave003
etc...
Finally, and this is also a bit tedious, via SuperPutty make every box SSH to each box in turn and check that all’s well. Ie, send them all:
hduser@node:~$ ssh master
…check that they all have…
hduser@node:~$ ssh slave001
… check that they all have… etc.
This is a one-time thing; after any box has connected to any other one time, the link between them remains.
Java
The next prerequisite to sort is a Java environment, as the Hadoop core is written Java (although you can harness the power of MapReduce in any language you please, as we shall see). If you’re fortunate, your machines will have internet access, in which case fire the following command to them all using SuperPutty:
hduser@node:~$ sudo apt-get install openjdk-6-jre
If like mine, however, your machines were considered ticking chemical time bombs by infrastructure and hence weren’t granted internet access, what you’ll want to do is download a JDK to a computer that does have internet access and can also see your Hadoop boxes on the network, and fire the files over from there. So on your internet-connected box:
32 bit version:
hduser@node:~$ wget --no-cookies --no-check-certificate --header "Cookie: gpw_e24=http://www.oracle.com/" http://download.oracle.com/otn-pub/java/jdk/6u34-b04/jre-6u34-linux-i586.bin
64 bit version:
hduser@node:~$ wget --no-cookies --no-check-certificate --header "Cookie: gpw_e24=http://www.oracle.com/" http://download.oracle.com/otn-pub/java/jdk/6u45-b06/jdk-6u45-linux-x64.bin
Now then! Each of your Hadoop nodes wants to connect to this box and pull over the Java files. Find its IP by typing ifconfig, and then fire this command to all of your Hadoop nodes:
hduser@node:~$ scp user@internetbox:/locationoffile/rightarchtecturefile.bin $HOME
Be careful to get the edition matching the machine, be it 32bit or 64bit.
Now execute the following on the Hadoop machines to install Java…
32 bit machines:
hduser@node:~$ chmod u+x jre-6u34-linux-i586.bin
hduser@node:~$ ./jre-6u34-linux-i586.bin
hduser@node:~$ sudo mkdir -p /usr/lib/jvm
hduser@node:~$ sudo mv jre1.6.0_34 /usr/lib/jvm/
hduser@node:~$ sudo update-alternatives --install "/usr/bin/java" "java" "/usr/lib/jvm/jre1.6.0_34/bin/java" 1
hduser@node:~$ sudo update-alternatives --install "/usr/lib/mozilla/plugins/libjavaplugin.so" "mozilla-javaplugin.so" "/usr/lib/jvm/jre1.6.0_34/lib/i386/libnpjp2.so" 1
hduser@node:~$ sudo update-alternatives --install "/usr/bin/javaws" "javaws" "/usr/lib/jvm/jre1.6.0_34/bin/javaws" 1
hduser@node:~$ sudo update-alternatives --config java
hduser@node:~$ sudo update-alternatives --config javac
hduser@node:~$ export JAVA_HOME=/usr/lib/jvm/jre1.6.0_34/
64 bit machines:
hduser@node:~$ chmod u+x jdk-6u45-linux-x64.bin
hduser@node:~$ ./jdk-6u45-linux-x64.bin
hduser@node:~$ sudo mv jdk1.6.0_45 /opt
hduser@node:~$ sudo update-alternatives --install "/usr/bin/java" "java" "/opt/jdk1.6.0_45/bin/java" 1
hduser@node:~$ sudo update-alternatives --install "/usr/bin/javac" "javac" "/opt/jdk1.6.0_45/bin/javac" 1
hduser@node:~$ sudo update-alternatives --install "/usr/lib/mozilla/plugins/libjavaplugin.so" "mozilla-javaplugin.so" "/opt/jdk1.6.0_45/jre/lib/amd64/libnpjp2.so" 1
hduser@node:~$ sudo update-alternatives --install "/usr/bin/javaws" "javaws" "/opt/jdk1.6.0_45/bin/javaws" 1
hduser@node:~$ sudo update-alternatives --config java
hduser@node:~$ sudo update-alternatives --config javac
hduser@node:~$ export JAVA_HOME=/opt/jdk1.6.0_45/
Finally, test by firing all machines
hduser@node:~$ java --version
You should see something like this:
hduser@node:~$ java -version
java version "1.6.0_45"
Java(TM) SE Runtime Environment (build 1.6.0_45-b06)
Java HotSpot(TM) 64-Bit Server VM (build 20.45-b01, mixed mode)
Installing Hadoop
Download Hadoop 2.2.0 into the directory /usr/local from the best possible source:
hduser@node:~$ cd /usr/local
hduser@node:~$ wget http://mirror.ox.ac.uk/sites/rsync.apache.org/hadoop/core/hadoop-2.2.0/hadoop-2.2.0.tar.gz
If your boxes don’t have internet connectivity, use the same workaround we used above to circuitously get Java.
Unzip, tidy up and make appropriate ownership changes:
hduser@node:~$ sudo tar xzf hadoop-2.2.0.tar.gz
hduser@node:~$ sudo mv hadoop-2.2.0 hadoop
hduser@node:~$ sudo chown -R hduser:hadoop hadoop
Finally, append the appropriate environment variable settings and aliases to the bash configuration file:
hduser@node:~$ echo "" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "export HADOOP_HOME=/usr/local/hadoop" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "" | sudo tee -a $HOME/.bashrc
#32 bit version:
hduser@node:~$ echo "export JAVA_HOME=/usr/lib/jvm/jre1.6.0_34" | sudo tee -a $HOME/.bashrc
#64 bit version:
hduser@node:~$ echo "export JAVA_HOME=/opt/jdk1.6.0_45" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "unalias fs &> /dev/null" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "alias fs &>"hadoop fs"" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "unalias hls &> /dev/null" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "alias hls="fs -ls" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "" | sudo tee -a $HOME/.bashrc
hduser@node:~$ echo "export PATH=$PATH:$HADOOP_HOME/bin" | sudo tee -a $HOME/.bashrc
There are a few changes that must be made to the configuration files in /usr/local/hadoop/etc/hadoop which inform the HDFS and MapReduce layers. Editing these on every machine at once via SuperPutty requires skill, especially when, having made the changes, you realise that you can’t send an “escape” character to every machine at once. There’s a solution involving mapping other, sendable, characters to the escape key, but that’s “out of scope” here 😉 Here’s what the files should look like.
core-site.xml
It needs to look like this on all machines, master and slave alike:
[code language=”xml”]
<configuration>
<property>
<name>fs.default.name</name>
<value>hdfs://master:9000</value>
</property>
<property>
<name>hadoop.tmp.dir</name>
<value>/usr/local/hadoop/tmp</value>
</property>
</configuration>[/code]
hadoop-env.sh
There’s only one change that needs to be made to this mofo; locate the line which specifies JAVA_HOME (helpfully commented with “the Java implementation to use”). Assuming a Java setup like that described above, this should read
32 bit machines:
export JAVA_HOME=/usr/lib/jvm/jre1.6.0_34/
64 bit machines:
export JAVA_HOME=/opt/jdk1.6.0_45/
hdfs-site.xml
This specifies the replication level of file blocks. Note that your physical storage size will be divided by this number to give the storage you’ll have in HDFS.
[code language=”xml”]
<configuration>
<property>
<name>dfs.replication</name>
<value>3</value>
</property>
</configuration>[/code]
Additionally, it’s necessary to create a local directory on each box for Hadoop to use:
hduser@node:~$ sudo mkdir -p /app/hadoop/tmp
hduser@node:~$ sudo chown hduser:hadoop /app/hadoop/tmp
mapred-site.xml
Which MapReduce implementation to use. At the moment we’re on YARN (“Yet Another Resource Negotiator”…………).
[code language=”xml”]
<configuration>
<property>
<name>mapreduce.framework.name</name>
<value>yarn</value>
</property>
</configuration>[/code]
yarn-site.xml
Controls the actual MapReduce configuration. Without further ado, this is what you want:
[code language=”xml”]
<configuration>
<property>
<name>yarn.nodemanager.aux-services</name>
<value>mapreduce_shuffle</value>
</property>
<property>
<name>yarn.nodemanager.aux-services.mapreduce_shuffle.class</name>
<value>org.apache.hadoop.mapred.ShuffleHandler</value>
</property>
<property>
<name>yarn.resourcemanager.resource-tracker.address</name>
<value>master:8025</value>
</property>
<property>
<name>yarn.resourcemanager.scheduler.address</name>
<value>master:8030</value>
</property>
<property>
<name>yarn.resourcemanager.address</name>
<value>master:8040</value>
</property>
</configuration>[/code]
Slaves
In short, the master needs to consider itself and every other node a slave. Each slave needs to consider itself, and itself only, a slave. The entirety of your slaves file ought to look like this:
Master:
master
slave001
slave002
slave003
etc
Slave xyz:
slavexyz
Formatting the Filesystem
Much like manually deleting your data, formatting a HDFS filesystem containing data will delete any data you might have in it, so don’t do that if you don’t want to delete your data. Warnings notwithstanding, execute the following on the master node to format the HDFS namespace:
hduser@master:~$ cd /usr/local/hadoop
hduser@master:~$ bin/hadoop namenode -format
Bringing up the Cluster
This is the moment that the band strikes up. If you’re not already there, switch to the Hadoop directory…
hduser@master:~$ cd /usr/local/hadoop
Fire this shizz to start the DFS layer:
hduser@master:/usr/bin/hadoop$ sbin/start-dfs.sh
You should see this kind of thing:
WARN util.NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
starting namenodes on [master]
master: starting namenode, logging to /usr/local/hadoop/logs/hadoop-hduser-namenode-master.out
slave001: starting datanode, logging to /usr/local/hadoop/logs/hadoop-hduser-datanode-slave001.out
slave002: starting datanode, logging to /usr/local/hadoop/logs/hadoop-hduser-datanode-slave002.out
slave003: starting datanode, logging to /usr/local/hadoop/logs/hadoop-hduser-datanode-slave003.out
...etc
Now start the MapReduce layer:
hduser@master:/usr/local/hadoop$ sbin/start-yarn.sh
Expect to be greeted by something like this:
starting yarn daemons
starting resourcemanager, logging to /usr/local/hadoop/logs/yarn-hduser-resourcemanager-master.out
slave001: starting nodemanager, logging to /usr/local/hadoop/logs/yarn-hduser-nodemanager-slave001.out
slave002: starting nodemanager, logging to /usr/local/hadoop/logs/yarn-hduser-nodemanager-slave002.out
slave003: starting nodemanager, logging to /usr/local/hadoop/logs/yarn-hduser/nodemanager-slave003.out
...
Also start the job history server…
hduser@master:/usr/local/hadoop$ sbin/mr-jobhistory-daemon.sh start historyserver
Surveying One’s Empire
By this stage your Hadoop cluster is humming like a dynamo. There are several web interfaces which provide a tangible window into the specification of the cluster as a whole…
For the DFS layer, have a look at http://master:50070.
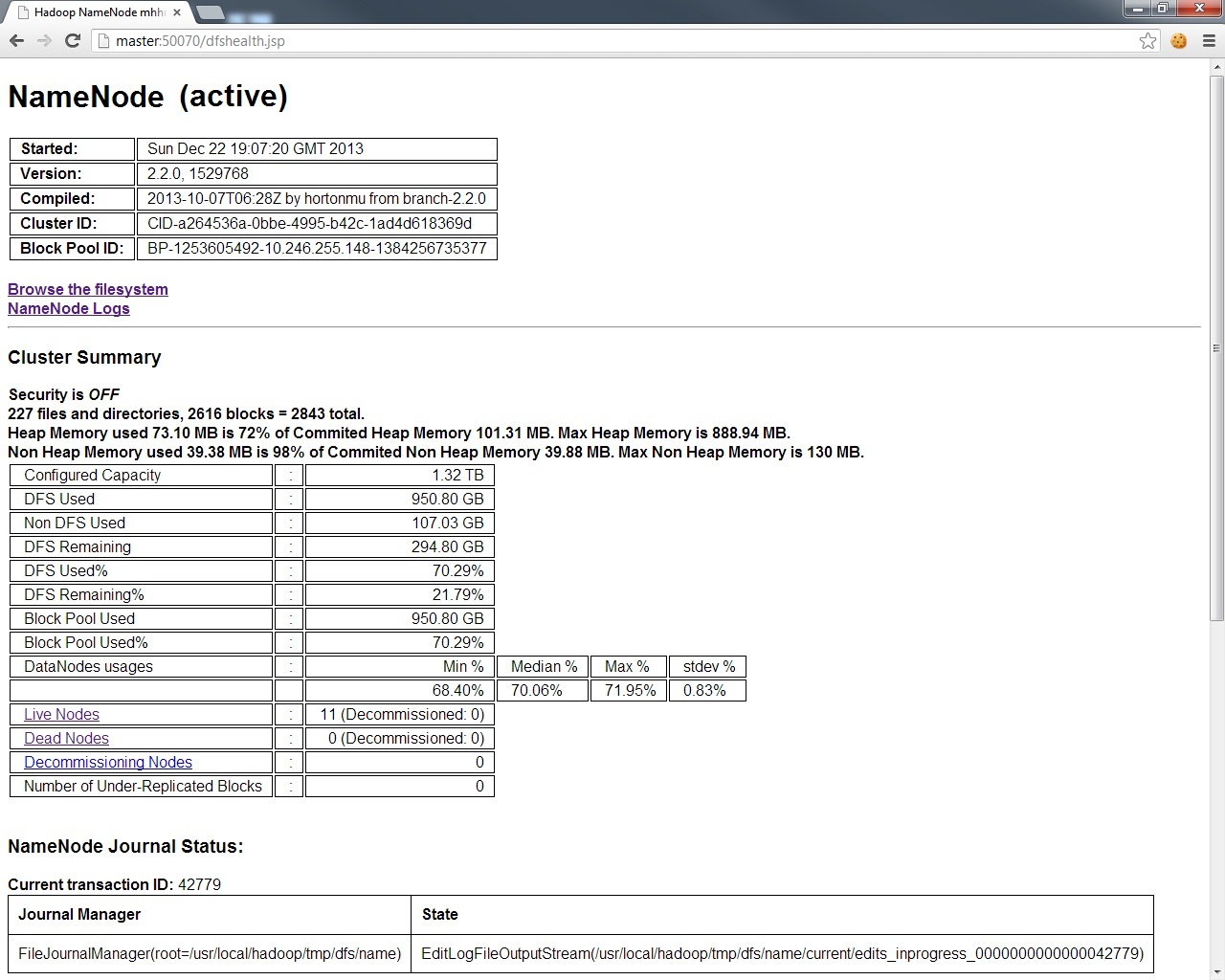
And for a breakdown of the exact condition of each node in your DFS layer,
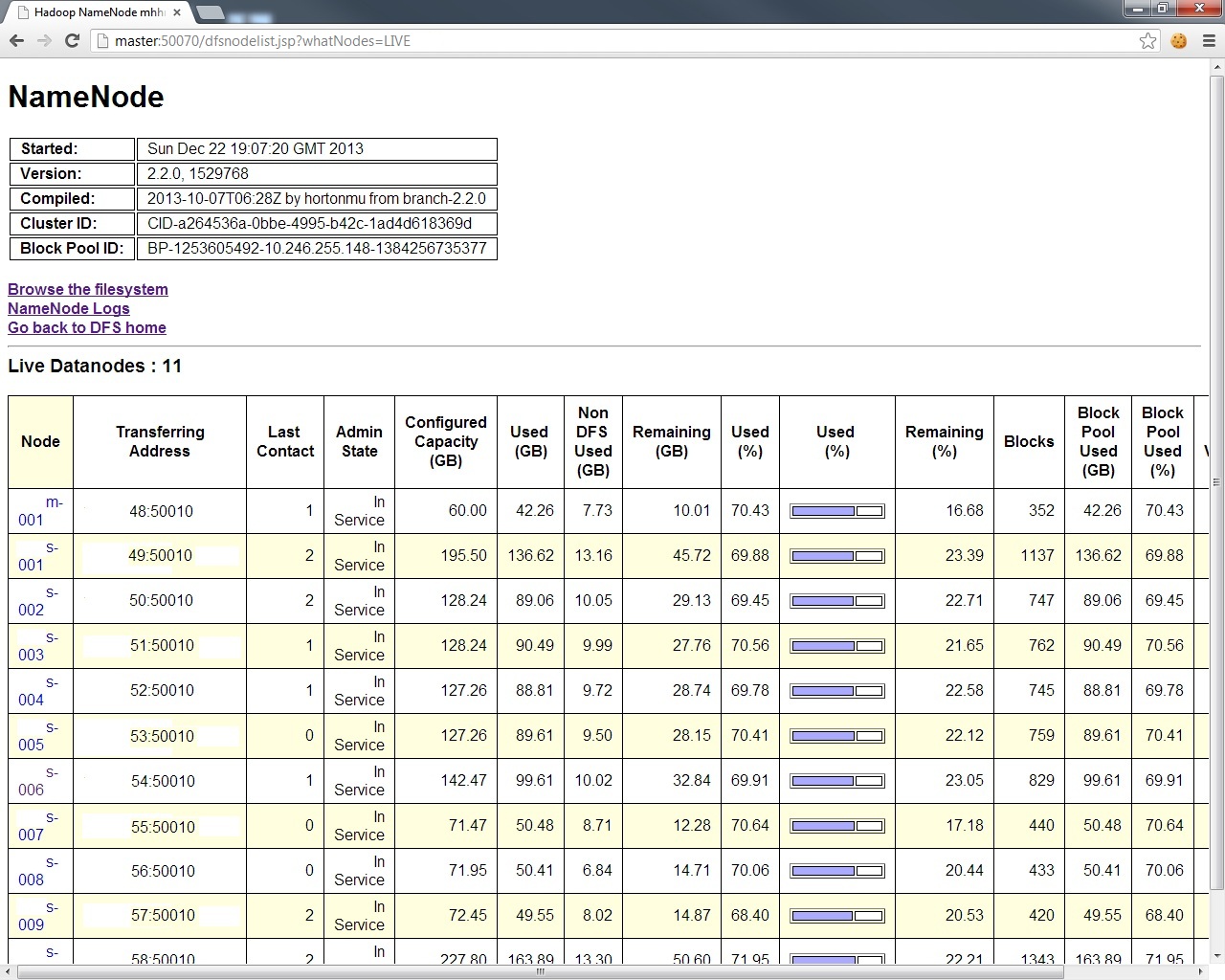
And for the MapReduce layer, look at http://master:8088,
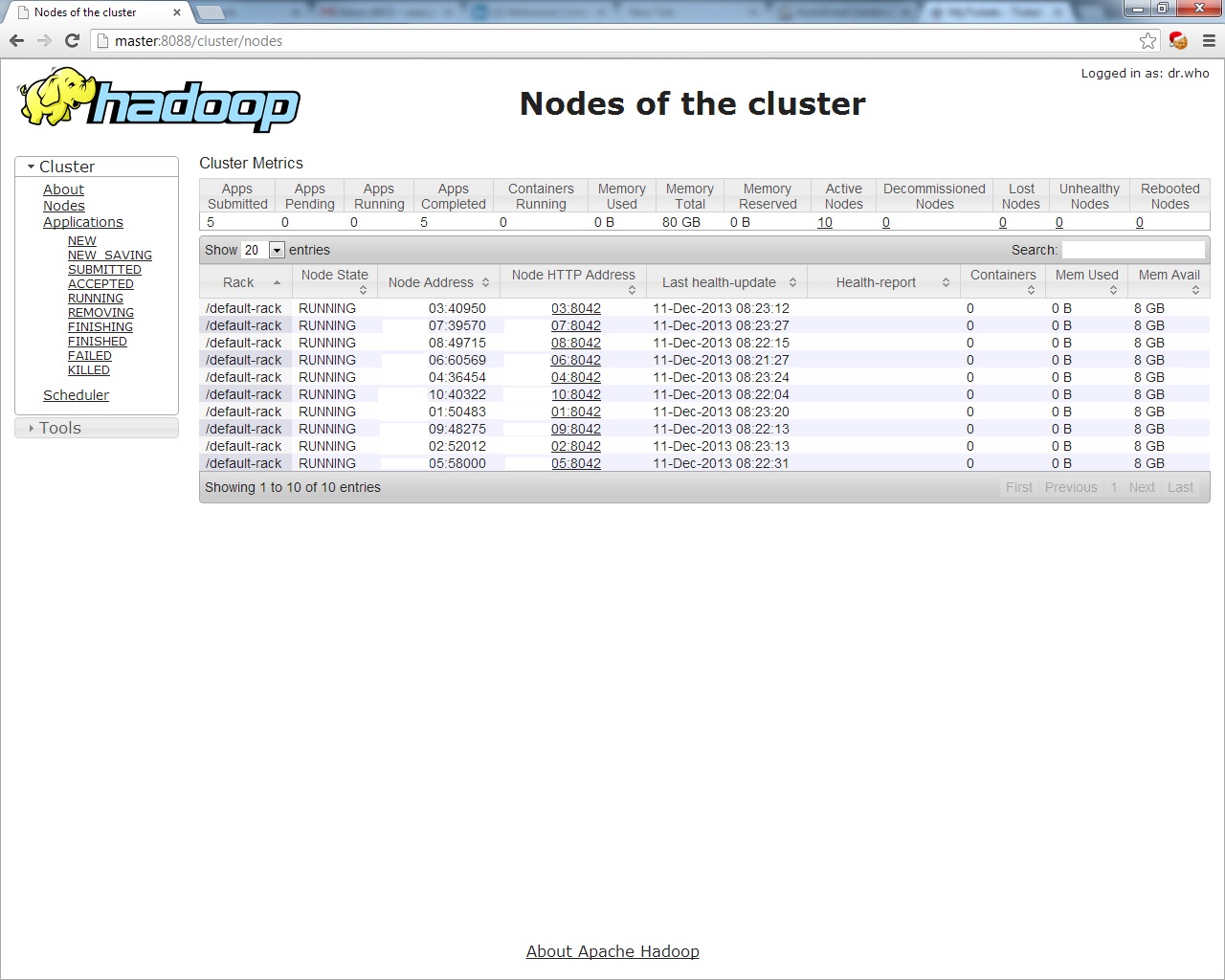
The First Distributed MapReduce
MapReduce is nothing more than a certain way to phrase a script to process a file, which is friendly to distributed computing. There’s a mapper, and a reducer. The “mapper” must be able to process any arbitrary fragment of the file (eg, count the number of occurrences of something within that fragment), independently and obliviously of the contents of the rest of the file. This is why it’s so scalable. The “reducer” aggregates the outputs of the mappers to give the final result (eg, sum up the occurrences of something reported by each of the mappers to give the total number of occurrences). Again, the way that you only have to write the mapper and reducer, and Hadoop handles the rest (deploying a copy of the mapper to every worker node, “shuffling” the mapper outputs for the reducer, re-allocating failed maps etc), is why Hadoop is well good. Indeed, a well-maintained cluster is much like American dance/rap duo LMFAO: every day it’s shuffling.
Later in this blog we’ll address how to write MapReduces; for now let’s just perform one and let the cluster stretch its legs for the first time.
Make a cheeky text file (example.txt):
Example text file
Contains example text
Make a directory in HDFS, lob the new file in there, and check that it’s there:
hduser@master:/usr/local/hadoop$ bin/hadoop fs -mkdir /test
hduser@master:/usr/local/hadoop$ bin/hadoop fs -put example.txt /test
hduser@master:/usr/local/hadoop$ bin/hadoop fs -ls /test
WARN util.NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
Found 1 items
-rw-r--r-- 3 hduser supergroup 50 2013-12-23 09:09 /test/example.txt
As you can see, the Hadoop file system commands are very similar to the normal Linux ones. Now run the example MapReduce:
hduser@master:/usr/local/hadoop$ bin/hadoop jar share/hadoop/mapreduce/hadoop-mapreduce-examples-2.2.0.jar wordcount /test /testout
Hadoop will immediately inform you that a load of things are now deprecated – ignore these warnings, it seems that deprecation is the final stage in creating new Hadoop modules – and then more interestingly keep you posted on the progress of the job…
INFO mapreduce.JobSubmitter: Submitting tokens for job: job_1387739551023_0001
INFO impl.YarnClientImpl: Submitted application application_1387739551023_0001 to ResourceManager at master/1.1.1.1:8040
INFO mapreduce.Job: The url to track the job: http://master:8088/proxy/application_1387739551023_0001/
INFO mapreduce.Job: Running job: job_1387739551023_0001
INFO mapreduce.Job: Job job_1387739551023_0001 running in uber mode : false
INFO mapreduce.Job: map 0% reduce 0%
INFO mapreduce.Job: map 100% reduce 0%
INFO mapreduce.Job: map 100% reduce 100%
INFO mapreduce.Job: Job job_1387739551023_0001 completed successfully
INFO mapreduce.Job: Counters: 43
File System Counters
FILE: Number of bytes read=173
FILE: Number of bytes written=158211
FILE: Number of read operations=0
FILE: Number of large read operations=0
FILE: Number of write operations=0
HDFS: Number of bytes read=202
HDFS: Number of bytes written=123
HDFS: Number of read operations=6
HDFS: Number of large read operations=0
HDFS: Number of write operations=2
Job Counters
Launched map tasks=1
Launched reduce tasks=1
Rack-local map tasks=1
Total time spent by all maps in occupied slots (ms)=7683
Total time spent by all reduces in occupied slots (ms)=11281
Map-Reduce Framework
Map input records=2
Map output records=11
Map output bytes=145
Map output materialized bytes=173
Input split bytes=101
Combine input records=11
Combine output records=11
Reduce input groups=11
Reduce shuffle bytes=173
Reduce input records=11
Reduce output records=11
Spilled Records=22
Shuffled Maps =1
Failed Shuffles=0
Merged Map outputs=1
GC time elapsed (ms)=127
CPU time spent (ms)=2570
Physical memory (bytes) snapshot=291241984
Virtual memory (bytes) snapshot=1030144000
Total committed heap usage (bytes)=181075968
Shuffle Errors
BAD_ID=0
CONNECTION=0
IO_ERROR=0
WRONG_LENGTH=0
WRONG_MAP=0
WRONG_REDUCE=0
File Input Format Counters
Bytes Read=101
File Output Format Counters
Bytes Written=123
hduser@master:/usr/local/hadoop$
GLORY. We can examine the output thus:
hduser@master:/usr/local/hadoop$ bin/hadoop fs -ls /testout
WARN util.NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
Found 2 items
-rw-r--r-- 3 hduser supergroup 50 2013-12-23 09:10 /testout/_SUCCESS
-rw-r--r-- 3 hduser supergroup 50 2013-12-23 09:10 /testout/part-r-00000
hduser@master:/usr/local/hadoop$ bin/hadoop fs -cat /testout/part-r-00000
WARN util.NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
Contains 1
Example 1
example 1
file 1
text 2
Business value, delivered. If you want to retrieve the output file from HDFS back to your local filesystem, run
hduser@master:/usr/local/hadoop$ bin/hadoop fs -get /testout
hduser@master:/usr/local/hadoop$ ls | grep testout
testout
And there it is! Now that your Hadoop cluster is essentially a self-aware beacon of supercomputing, stay tuned for further posts on using Hadoop to do interesting/lucrative things! 🙂